 Weak mismatch elevates chances of amplification and so add strong mismatch to avoid false-amplification near the 3’-OH end of the primer (at -2 position ideally). You may wonder why we need to add a mismatch? The mismatch is the key factor in achieving the amplification. We have to design a forward primer in such a manner that for the normal allele the primer contains C (complementary to G) at 3’ end and the mutant primer contains T in place of C.Īlthough it works magically when we add a mismatched base near our SNP at the 3’ end. See the image above first, Suppose our DNA sequence has G-A point mutation viz, G in normal allele and A in place of G in the mutant allele. A few points to keep in mind while preparing are, Remember the primer must be allele-specific. Primer preparation and designing play a crucial role here. Results interpretations Primer designing:.We can divide the whole process into 4 separate steps: Hazardous radiolabeled probes aren’t involved here. The image represents the concept of a mismatch for ARMS-PCR. See the image below so you can understand the concept easily. Why aren’t we using high-fidelity Taq DNA polymerase in ARMS PCR? Think about this question, I will answer it later. Mismatch indeed alters the annealing temperature for different sets of primers. Here the mismatch between the primer and the template DNA plays a crucial role in achieving the amplification. The mismatch concept:īy introducing a mismatch in a primer makes refractory amplification possible. Gel electrophoresis helps to examine results. Only a single set of primers can only amplify a specific allele present in a sample, during the reaction. To do this, researchers modify a few bases from the primer’s 3’ OH end. The 3’ end of the primer is modified in such a way that one primer can amplify a mutant allele while the other can amplify the normal allele. The principle of the present technique relies on the modification of primers to amplify a specific allele. Henceforth, known as Amplification Refractory Mutation System. The mutant allele-specific primers are refractory (resistant) to normal allele and vice versa. Two different primer sets are prepared for separate alleles. While gel electrophoresis prepares results to evaluate. PCR amplifies each allele simultaneously, afterward. However, we need to do a minor change in primers to amplify each allele. Each set of specific primers is designed for each specific allele. Meaning, If you wish to amplify only a mutant allele, design a primer set accordingly and amplify it using this technique. The Allele-specific PCR has the power to detect a single specific allele. When a mutation occurs, it produces two different alleles for a gene one mutant one and another normal one. Wrapping up: What is ARMS or Allele-Specific PCR?Īlleles are alternative gene forms.This article will boost your PCR knowledge and will surely help you in your study. I have also explained the advantages, disadvantages and applications. The present post comprises information on ARMS- PCR, its principle, process, protocol and steps. That’s all we are going to discuss in the present article. The present technique has applications in detecting SNPs (Single nucleotide polymorphism) and genotyping. Newton and coworkers discovered the ARMS-PCR or allele-specific PCR technique. After a few years of the discovery of the actual PCR technique, C. Kary Mullis described the technique of in vitro amplification in the year 1983. It uses the word “Allele” many times, the reason is its specific amplification capacity for a specific allele or DNA sequence. 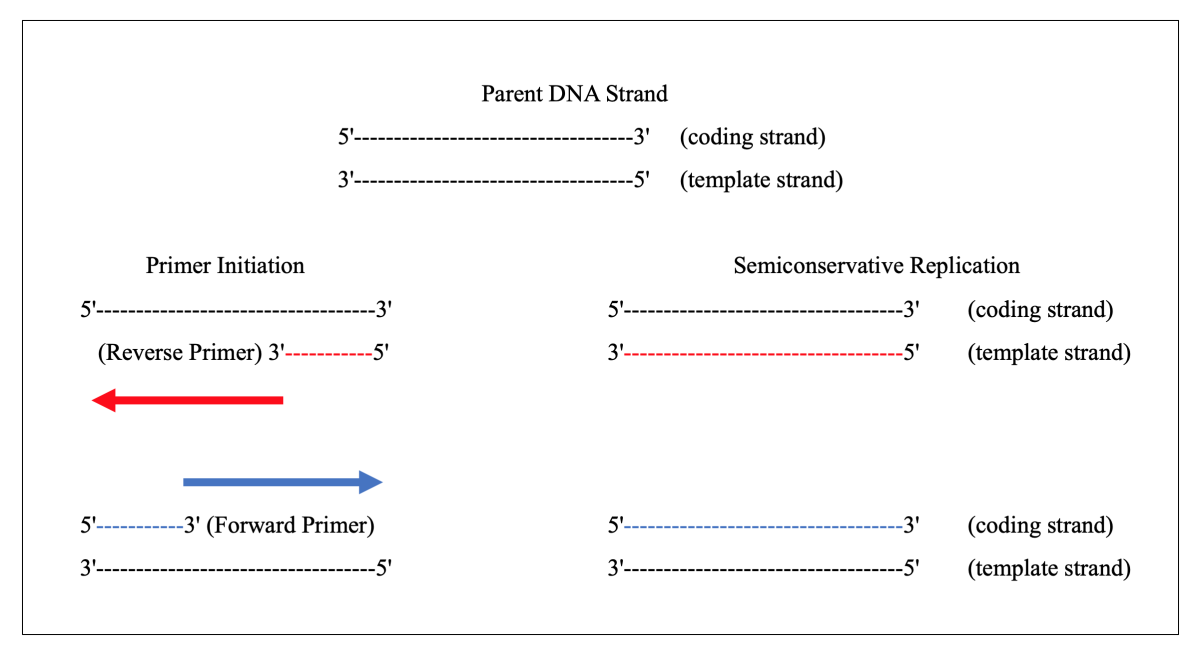
PASA acronyms as PCR Amplification of Specific Allele and AS- PCR stand for Allele- Specific PCR. Let us know the full names, ARMS acronym as Amplification Refractory Mutation System. It has other names like allele-specific PCR, PASA or AS- PCR, all have similar applications. Many variations of PCR exist depending upon the assay requirement, ARMS-PCR is one among them. It’s a temperature-dependent amplification technique that relies on Taq DNA polymerase. The PCR technique can detect mutations like deletion, duplications, insertion or single base change- SNP. “A method used to detect a single base change or SNP using sequence-specific primers is referred to as ARMS or Allele-specific PCR.” 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
